"Twins" retrives similar metabolites in chemical structure.
|
|
"Twins" retrives similar metabolites in chemical structure. |
| last update : 2020.01.06 |
| INPUT WORD = C00039813 , 50% or more |
|
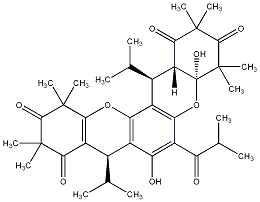
| [ Metabolite Name : Myrtucommulone D , (+)-Myrtucommulone D ] | |
| Number of matched data : 52 | |
| CID | Metabolite Name | Structural formula | Metabolite Activity | Similarity (%) |
|---|---|---|---|---|
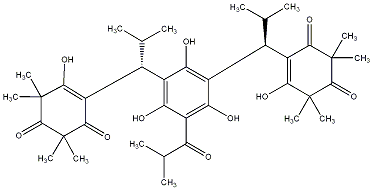
| C00039812 | Myrtucommulone A (+)-Myrtucommulone A |
 |
93.88 | C00039815 | Myrtucommulone G (+)-Myrtucommulone G |
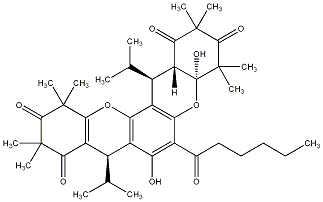 |
93.14 | C00039817 | Myrtucommulone I (+)-Myrtucommulone I |
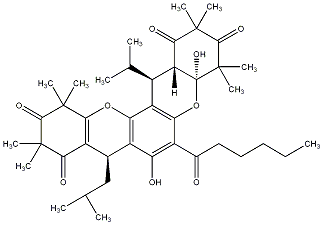 |
92.31 | C00039814 | Myrtucommulone F (+)-Myrtucommulone F |
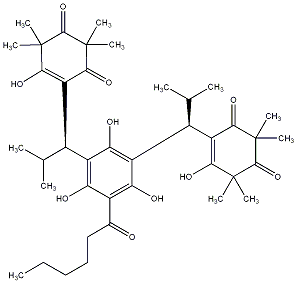 |
88.24 | C00039816 | Myrtucommulone H (+)-Myrtucommulone H |
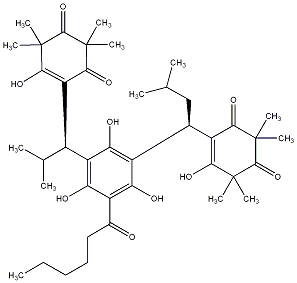 |
86.54 | C00002696 | Filixic acid BBB | 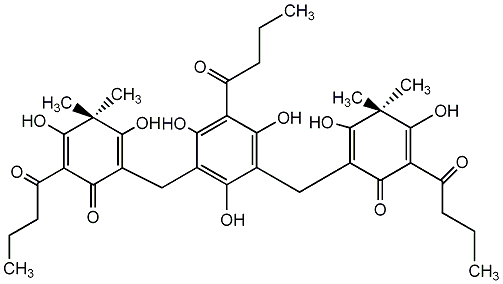 |
 |
82.65 | C00023931 | Hebelomic acid E | 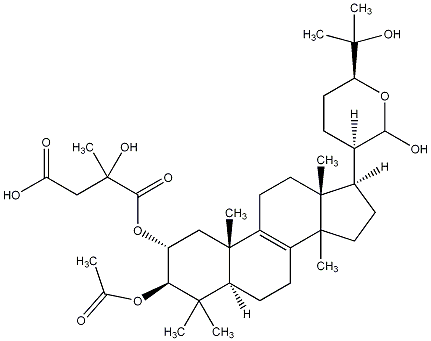 |
79.59 | C00032735 | Arjunetin 24-Deoxysericoside |
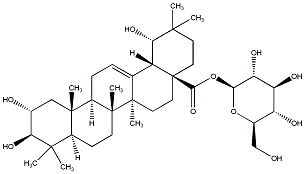 |
79.59 | C00037982 | Vernoguinoside | 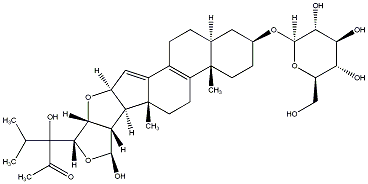 |
79.59 | C00032626 | 28-O-beta-D-glucopyranosyl-6beta,23-dihydroxytormentic acid | 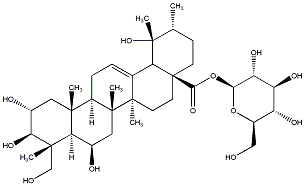 |
79.21 | C00031544 | 2alpha,3alpha,19alpha,24-Tetrahydroxyurs-12-en-28-oic acid 28-O-beta-D-glucopyranosyl ester (-)-4-Epi-niga-ichigoside F1 |
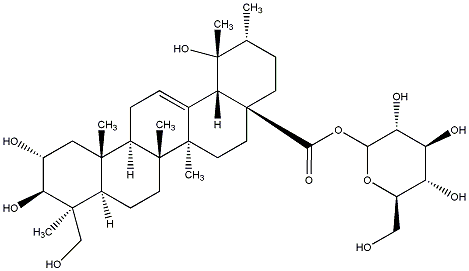 |
78.79 | C00032809 | Chebuloside II | 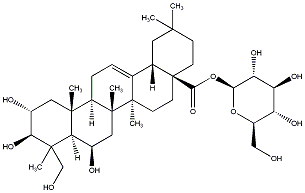 |
78.79 | C00032946 | CID is old! | 78.79 | C00033333 | Quadranoside VII (+)-Quadranoside VII |
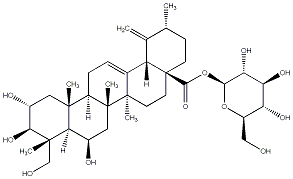 |
78.79 | C00033567 | 4-Epipinfaensin 4-epi-Pinfaensin |
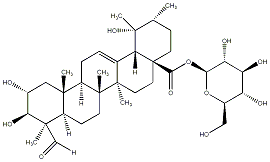 |
78.79 | C00034098 | Paradrymonoside (+)-Paradrymonoside |
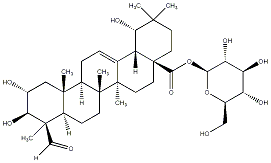 |
78.79 | C00035052 | Arjunglucoside I (+)-Arjunglucoside I |
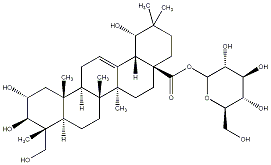 |
78.79 | C00035058 | Bellericoside (+)-Bellericoside |
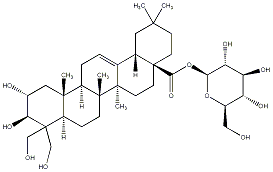 |
78.79 | C00035163 | Sericoside | 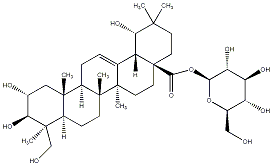 |
78.79 | C00039163 | Eryloside T (-)-Eryloside T |
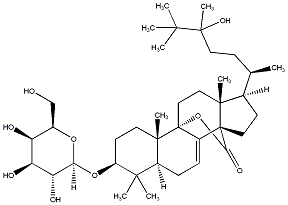 |
78.79 | C00040129 | Quadranoside III (+)-Quadranoside III |
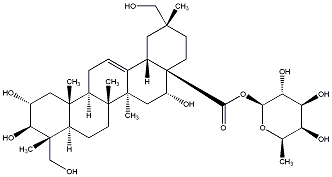 |
78.79 | C00047236 | Gambogenific acid (-)-Gambogenific acid |
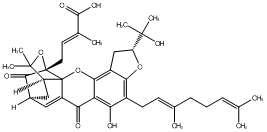 |
78.79 | C00048012 | Moluccensin C | 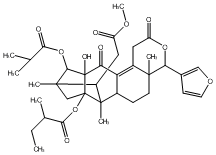 |
78.79 | C00048019 | Niga-ichigoside F1 (+)-Niga-ichigoside F1 |
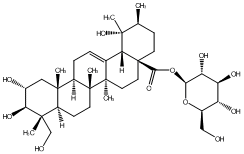 |
78.79 | C00048093 | Rubuside J (-)-Rubuside J |
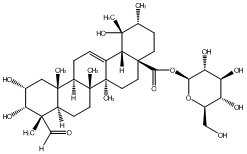 |
78.79 | C00047248 | Isogambogenin | 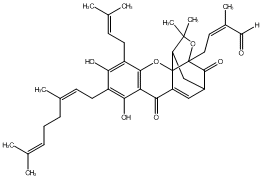 |
78.57 | C00044051 | 2-(beta-D-glucopyranosyloxy)-3,16,20-trihydroxy-9-methyl-19-norlanosta-5,24-diene-22-one | 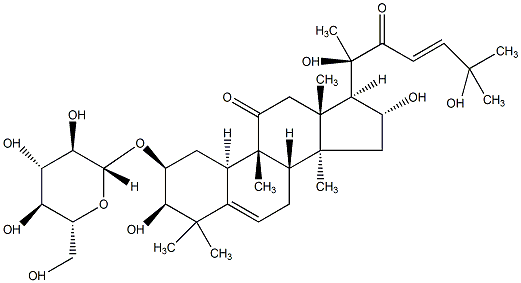 |
78.00 | C00033332 | Quadranoside VI (+)-Quadranoside VI |
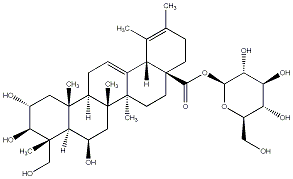 |
77.78 | C00048146 | Spirolide I | 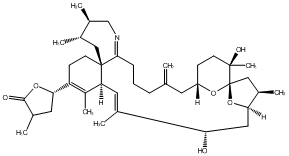 |
77.78 | C00040704 | Ximaolide C | 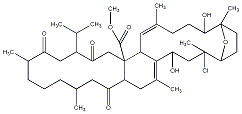 |
77.67 | C00030584 | Kalidiumoside D (-)-Kalidiumoside D |
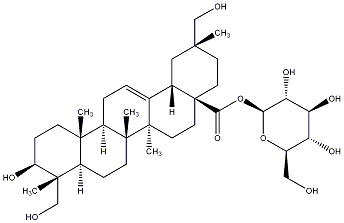 |
77.55 | C00030784 | Momordicoside G | 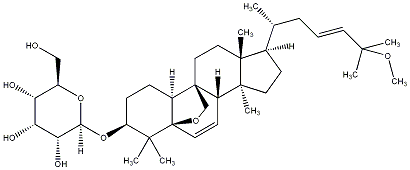 |
77.55 | C00032736 | Arjunglucoside II Arjunoglucoside II (+)-Arjunoglucoside II |
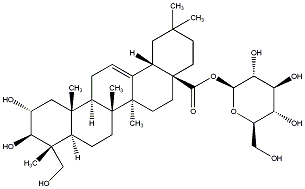 |
77.55 | C00032992 | Ginsenoside Rh6 (+)-Ginsenoside Rh6 |
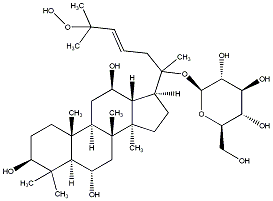 |
77.55 | C00033101 | Laetiposide F (+)-Laetiposide F |
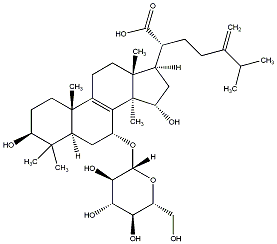 |
77.55 | C00033102 | Laetiposide G (+)-Laetiposide G |
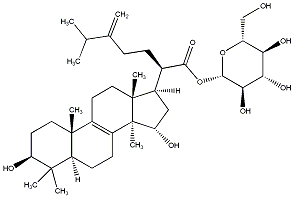 |
77.55 | C00038700 | Carbamidocyclophane E (+)-Carbamidocyclophane E |
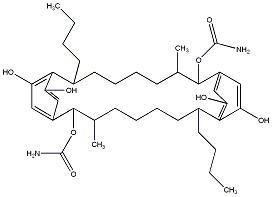 |
77.55 | C00039561 | Kolokoside A (-)-Kolokoside A |
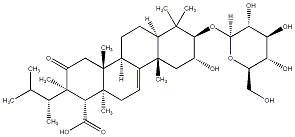 |
77.55 | C00039564 | Kolokoside D (-)-Kolokoside D |
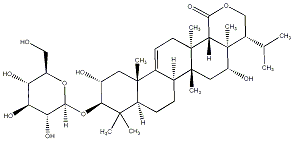 |
77.55 | C00045242 | Bryonioside A (+)-Bryonioside A |
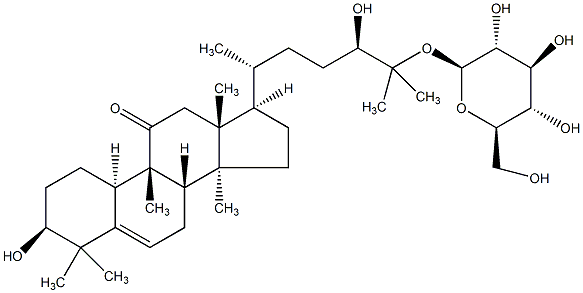 |
77.55 | C00045245 | Bryonioside D (+)-Bryonioside D |
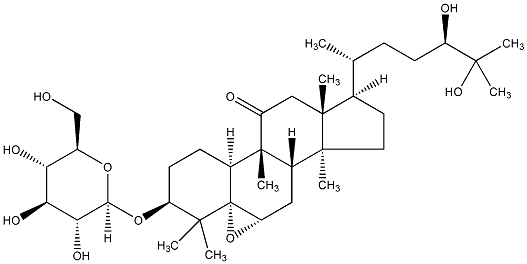 |
77.55 | C00046341 | Quadranoside IV (+)-Quadranoside IV |
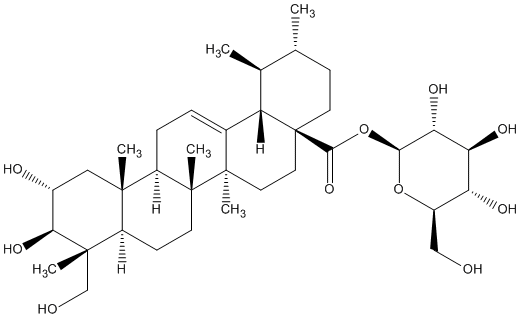 |
77.55 | C00046809 | Madlongiside C (+)-Madlongiside C |
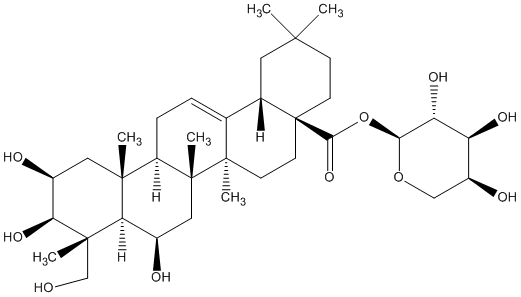 |
77.55 | C00048014 | Moluccensin F (+)-Moluccensin F |
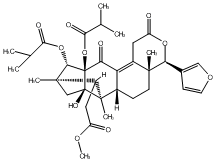 |
77.55 | C00048092 | Rubuside I (-)-Rubuside I |
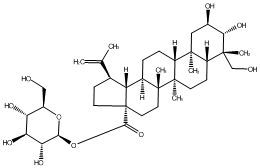 |
77.55 | C00049075 | 28-Glucosyl tormentate (+)-28-Glucosyl tormentate |
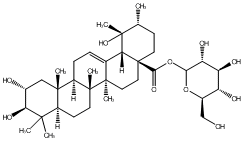 |
77.55 | C00039164 | Eryloside U (-)-Eryloside U |
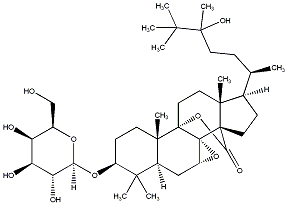 |
77.45 | C00014917 | 24-Demethylbafilomycin | 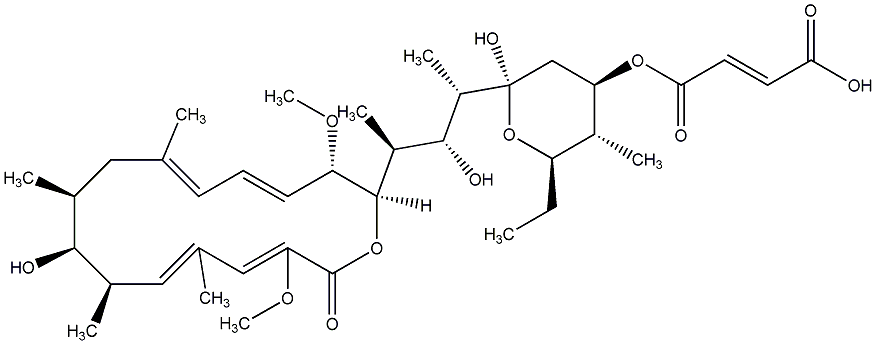 |
77.23 | C00032174 | Soulieoside C (-)-Soulieoside C |
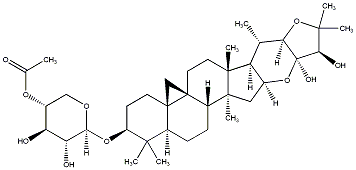 |
77.23 | C00032852 | Combreglucoside | 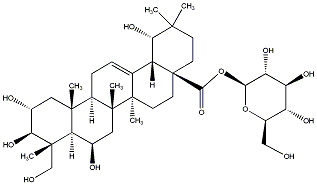 |
77.23 | C00041377 | Bisandrographolide B | 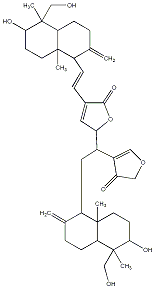 |
77.23 | C00041378 | Bisandrographolide C | 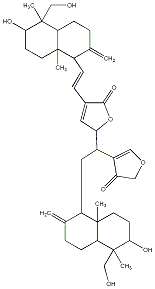 |
77.23 |